Validating the signal fidelity of novel soft neural probes
Filtering, feature extraction, clustering, regression and statistical testing of neural data.
This project, published in Nature Communications, aims to address the challenges of integrating soft neural probes with biological tissues. By utilizing high-resolution printing of liquid electronics, we can create interconnects directly on the cranium, minimizing mechanical mismatch and improving long-term stability.
As the sole statistician on the team, I applied signal processing and statistical testing to detect correlations between individual- and population-level neural responses to visual stimuli, thereby validating the signal fidelity of the novel neural probe. Additionally, I built an object-oriented MATLAB data pipeline (github.com/Jong-Min-Moon/neurosignal-matlab) for signal processing, feature extraction, and analysis of neural time-series data.
This page explains core concepts used in this project, including convolution, Fourier transform, Hilbert transform, and wavelet transform, and how they are used to analyze neural data in this paper.
References
computational neuroscience
-
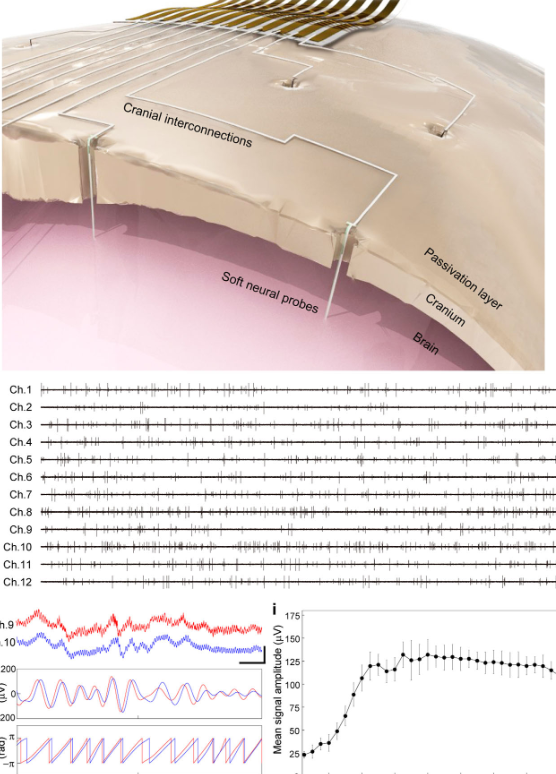 In-vivo integration of soft neural probes through high-resolution printing of liquid electronics on the craniumNature Communications, Feb 2024
In-vivo integration of soft neural probes through high-resolution printing of liquid electronics on the craniumNature Communications, Feb 2024
Project Updates
-
Data Analysis Detail: Mouse Behavioral Study
Details for the data analysis section of the Nature Communications paper 'In-vivo integration of soft neural probes through high-resolution printing of liquid electronics on the cranium'
-
Fundamentals of Phase Locking
Distinguishing Spike-Field Locking from Inter-Trial Phase Locking
-
Fundamentals of Neural Signal Decomposition
Spikes, LFPs, and frequency bands
-
Fundamentals of the Filter-Hilbert Method
Decoupling filtering from the analytic signal for better frequency control
-
Fundamentals of Continuous Wavelet Transform
Time-frequency decomposition using Analytic and Complex Morlet Wavelets
-
Fundamentals of Fourier Transform
From neural oscillations to frequency domain analysis
-
Fundamentals of Convolution
Inner product and convolution in signal processing and statistics
-
Fundamentals of Neural Recording
Non-invasive, invasive, intracellular, extracellular, and single-unit recording methods